A Culture of
Thinkers AND Doers
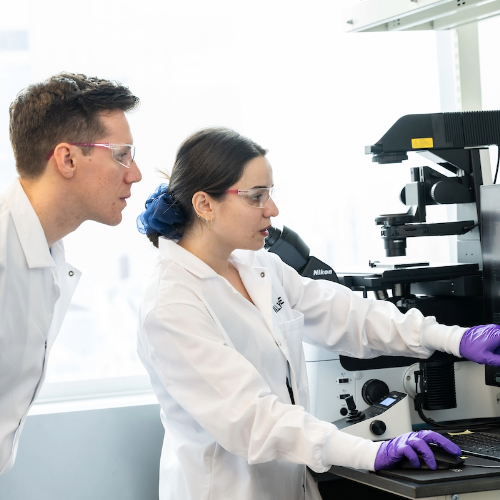








Our team brings together intrepid, dare-to-dream scientists with a group of highly accomplished and equally adventurous drug developers. The result is a rare combination of agile innovation and real-world focus — working together in service of truly important advances for human health.
As countless other biotechs have revealed, maintaining a balance between exploration and application isn’t easy. But it’s what separates great ideas from the ability to deliver real-world treatments and cures for patients. This is why we at Kallyope strive for a culture that champions scientific adventure and thrives on bringing together diverse perspectives from both inside and outside the company, leveraging some of the most accomplished academic scientists and external experts to help advance our work.
We are courageously curious. Profoundly connected. Made in New York City. And we always seek highly motivated individuals who wish to join our great adventure and help discover fundamentally new solutions to the world’s most challenging diseases. If that could be you, please email us at careers@kallyope.com.